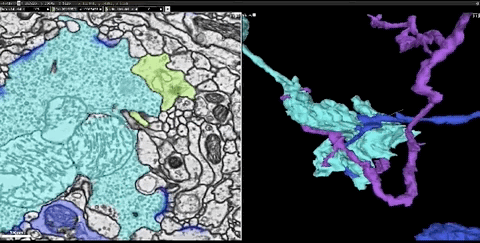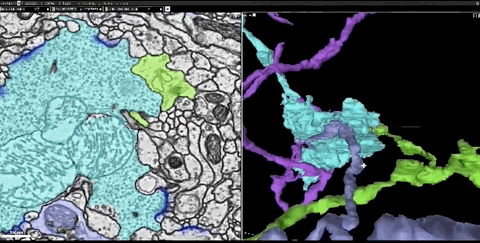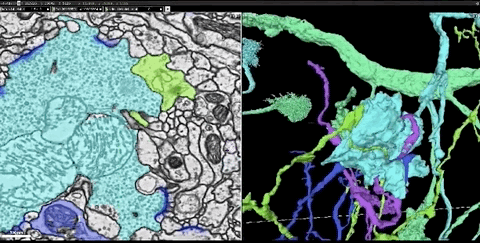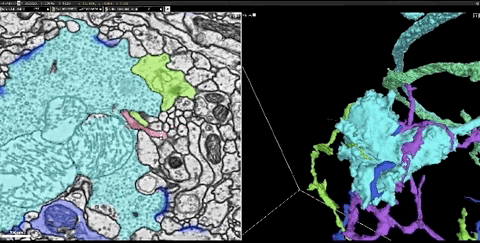
40 trillion pixels of complete Drosophila brain, automatically reconstructed.
Editor’s note: This article is from the WeChat public account “New Witness” ( ID: AI_era), edit Xiaoqin, Jin Lei, Zhang Jia.
Today, Google collaborated with the Howard Hughes Medical Institute (HHMI) and the University of Cambridge to launch an in-depth study of the fruit of the Drosophila brain – the brain that automatically rebuilds the entire fruit fly. They use thousands of Google Cloud TPUs to rebuild the entire fruit fly brain up to 40 trillion pixels. With a complete brain image, scientists are one step closer to understanding how the brain works.
Do you know? Drosophila is recognized as one of the most thoroughly studied organisms. Up to now, eight Nobel Prizes have been awarded to research using fruit flies, which have driven the development of molecular biology, genetics and neuroscience.
Scientists have always dreamed of understanding how the nervous system works by drawing the structure of a complete brain neural network.
One of the main goals of recent research is the brain of the fruit fly.
An important advantage of fruit flies is their size: Drosophila’s brain is relatively small, with only 100,000 neurons, compared to 100 million neurons in the brain of mice, the human brain There are 100 billion neurons.
This makes the brain of the fruit fly easier to study as a complete circuit.
Today, Google has teamed up with the Howard Hughes Medical Institute (HHMI) and the University of Cambridge to publish a new study of the brains of the Drosophila brain. Automatically reconstruct the brain of the entire fruit fly >.
Drosophila brain automatically rebuilds
This paper is entitled “Automatic reconstruction of continuous slice imaging of the Drosophila brain using the Flood-Filling network and local adjustments”:
A total of 16 researchers from Google, Howard Hughes Medical Research Institute (HHMI) Janelia Research Park and Cambridge University participated in the study. Among them, the first author Peter H. Li is a Google research scientist, the main research direction. Including general science, machine intelligence, machine perception.
Peter H. Li
They also provide a complete image of the Drosophila brain, which anyone can download and view, or browse online using interactive tools. They developed a 3D interactive interface called Neuroglancer.
This is not the first time that the Drosophila brain has been fully drawn. In January of this year, Science magazine covered the cover and introduced MIT and the Howard Hughes Medical Institute (HHMI) scientists have succeeded for fruit The entire brain of the fly was imaged and the resolution reached nanoscale. But that was still an artificial method, using two of the most advanced microscope techniques.
For decades, neuroscientists have been dreaming of drawing a complete map of the brain’s neural network, but for a human brain with 100 billion neural networks, the amount of data that needs to be processed is unimaginable. If you can automatically rebuild the Drosophila brain, it may be a step closer to automatically drawing the human brain.
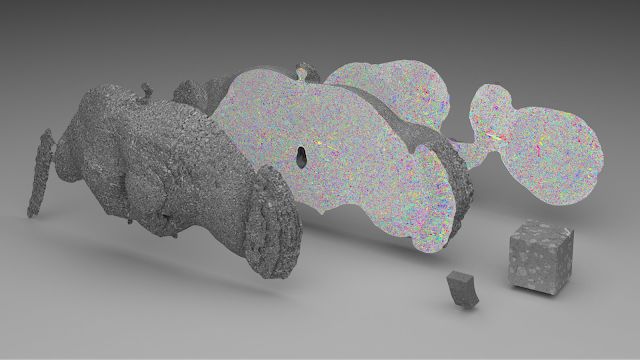
This is not the first time Peter H. Li’s team has attempted to map brain neurons using the AI method. They studied on smaller datasets in 2016 and 2018, respectively, as shown in the lower right corner of the image below.
a 40 trillion pixel 3D reconstruction of the Drosophila brain; the lower right corner is the smaller data set analyzed by Google AI in 2016 and 2018, respectively.
2018, Google collaborated with the Max Planck Institute for Neurobiology in Germany to develop a deep learning-based system that automatically maps neurons in the brain. They reconstructed a scanned image of the brain of a 1 million cubic micron zebra finch.
The researchers say that because of the high resolution of imaging, more than 1000 terabytes of brain tissue can produce more than 1000 terabytes of data. Therefore, this time rebuilding the brain of the whole fruit fly, I think the amount of data is huge.
It’s Google’s Cloud TPU for processing data, and it’s thousands!
Google AI director Jeff Dean also sighed on Twitter:
TPU really flies! GoogleAI scientists use TPU to Helps rebuild the nerve connections of the entire Drosophila brain!
Here, the new wisdom brings a detailed interpretation of this research:
40 trillion pixel fruit fly brain, automatically rebuilt!
In the course of the experiment, the main data set used was FAFB, which is an abbreviation for “full adult fly brain” (see the end of the relevant data set for related data).
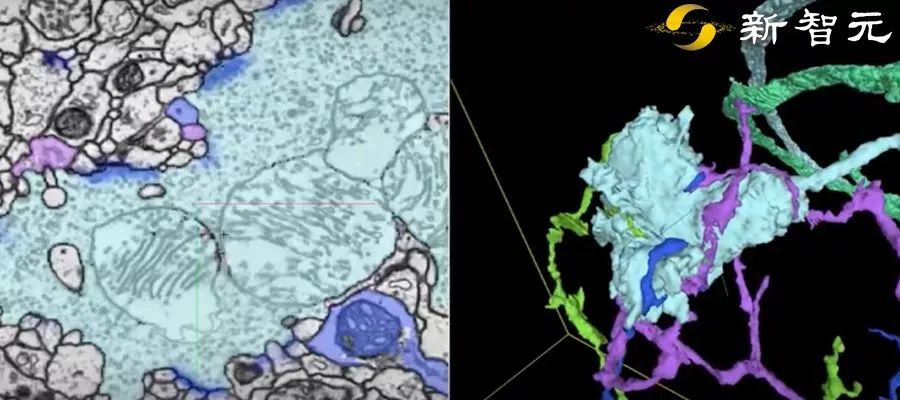
In this dataset, the researchers cut the fruit fly’s brain into thousands of ultra-thin sections of 40 nanometers, and then imaged each slice with a transmission electron microscope. This produces images of more than 40 trillion pixels of brain. And these 2D images are integrated into a coherent 3D fruit fly brain image.
Next, the researchers used thousands of cloud TPUs and applied Flood-Filling Network (FFN) to automate Track each neuron in the brain of the fruit fly.
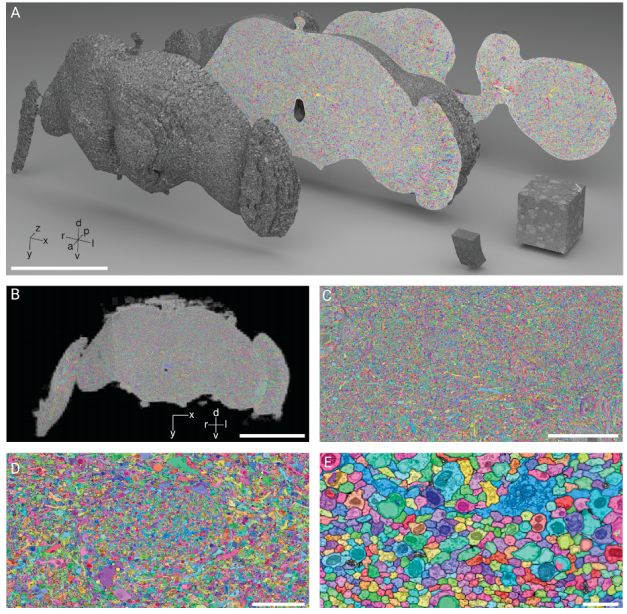
Dens segmentation of the entire Drosophila brain through FFN
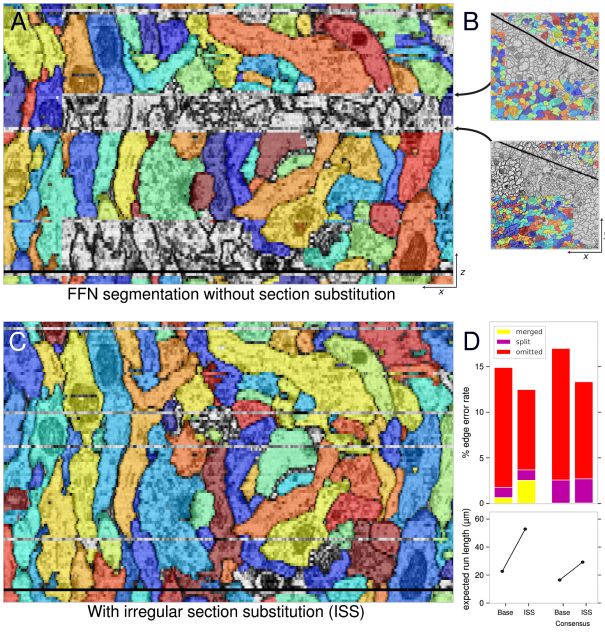
A in the above figure is a 3D rendered FAFB data set smoothed tissue mask. Any coronal slice (dataset XY plane) shows the entire internal FAFB-FFN1 segmentation. B-E shows the effect of increasing the zoom ratio.
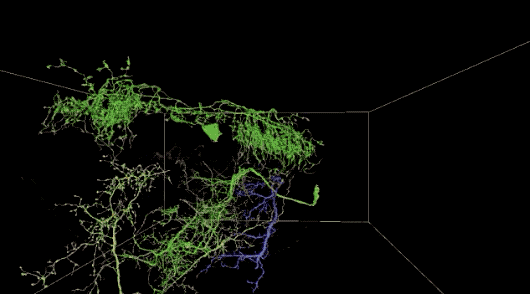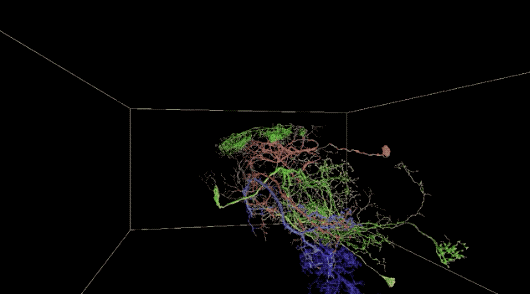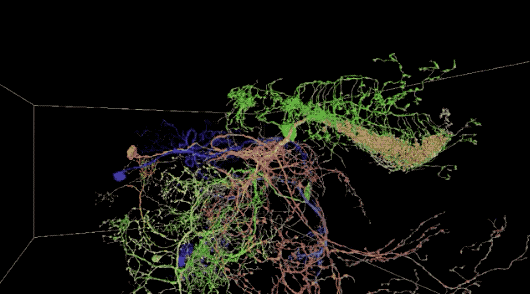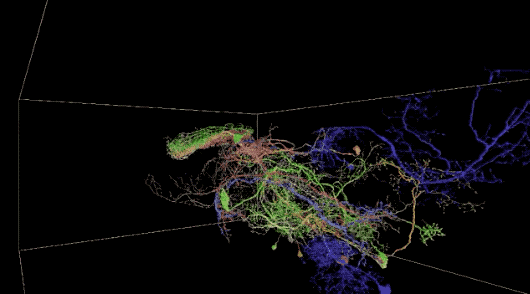
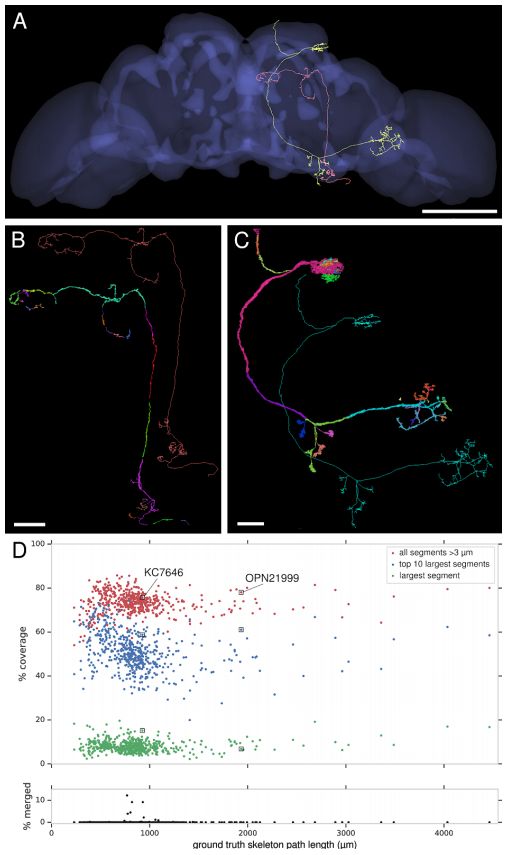
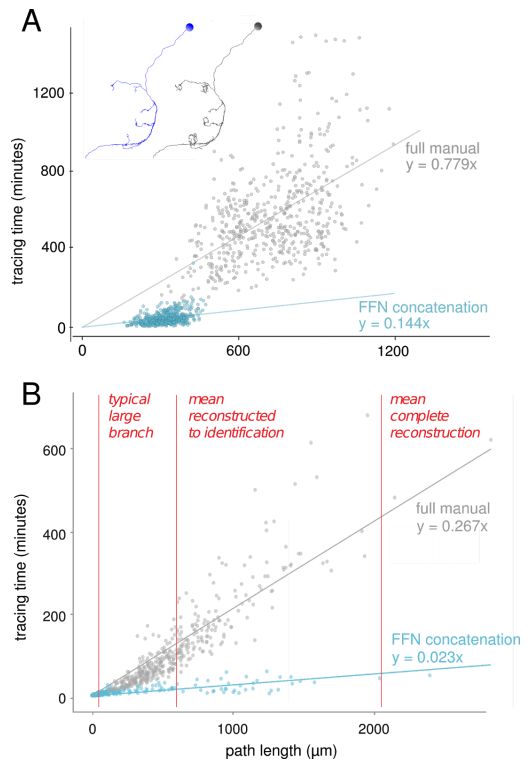
Automatic neuron reconstruction versus manual neuron tracking for verification
Although the overall performance of this algorithm is not bad, but when the alignment is not perfect (the image content in the continuous slice is unstable) or occasionally due to the loss of multiple serial slices during the imaging process, the performance will be decline.
To make up for this problem, the researchers combined FFN with two new programs.
First, the consistency between slices in each region in the 3D image is estimated, and then the content in the image is locally stabilized while the FFN tracks each neuron.
Secondly, the researchers used SECGAN to calculate missing slices in the image volume, and when using SECGAN, the researchers found that FFN can more reliably track the location of multiple missing slices.
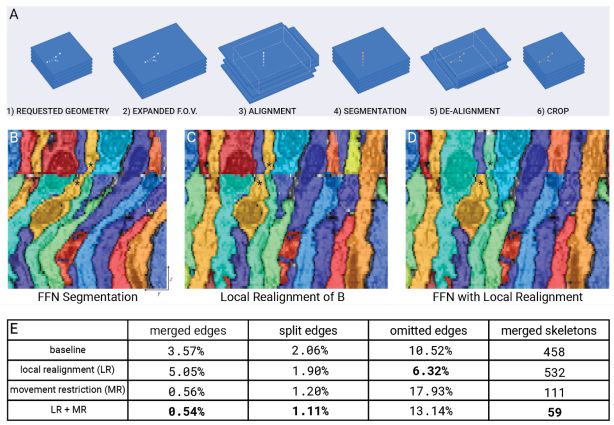
Local Realignment (LR)
Replacement of Irregular Sections
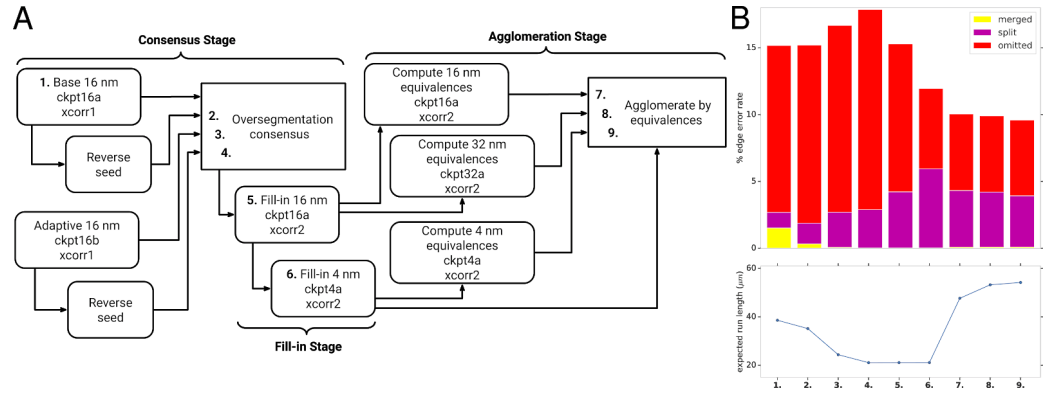
The split pipeline for the overall FAFB-FFN1
Segmentation-a